Project Introduction
Parcellations and neuroimaging atlases are ubiquitous in neuroimaging, namely because they allow for a principled reduction of features. This project focuses in particular on the question of choice of parcellation, in particular, how does choice of parcellation influence performance within a machine learning context (See Goals / Considerations for Machine Learning Based Neuroimaging). We perform a number of different experiments in order to probe this and related questions in detail.
This website acts as both a standalone project site and as online supplementary materials for the corresponding project paper - Why a website?
Base Experiment Setup
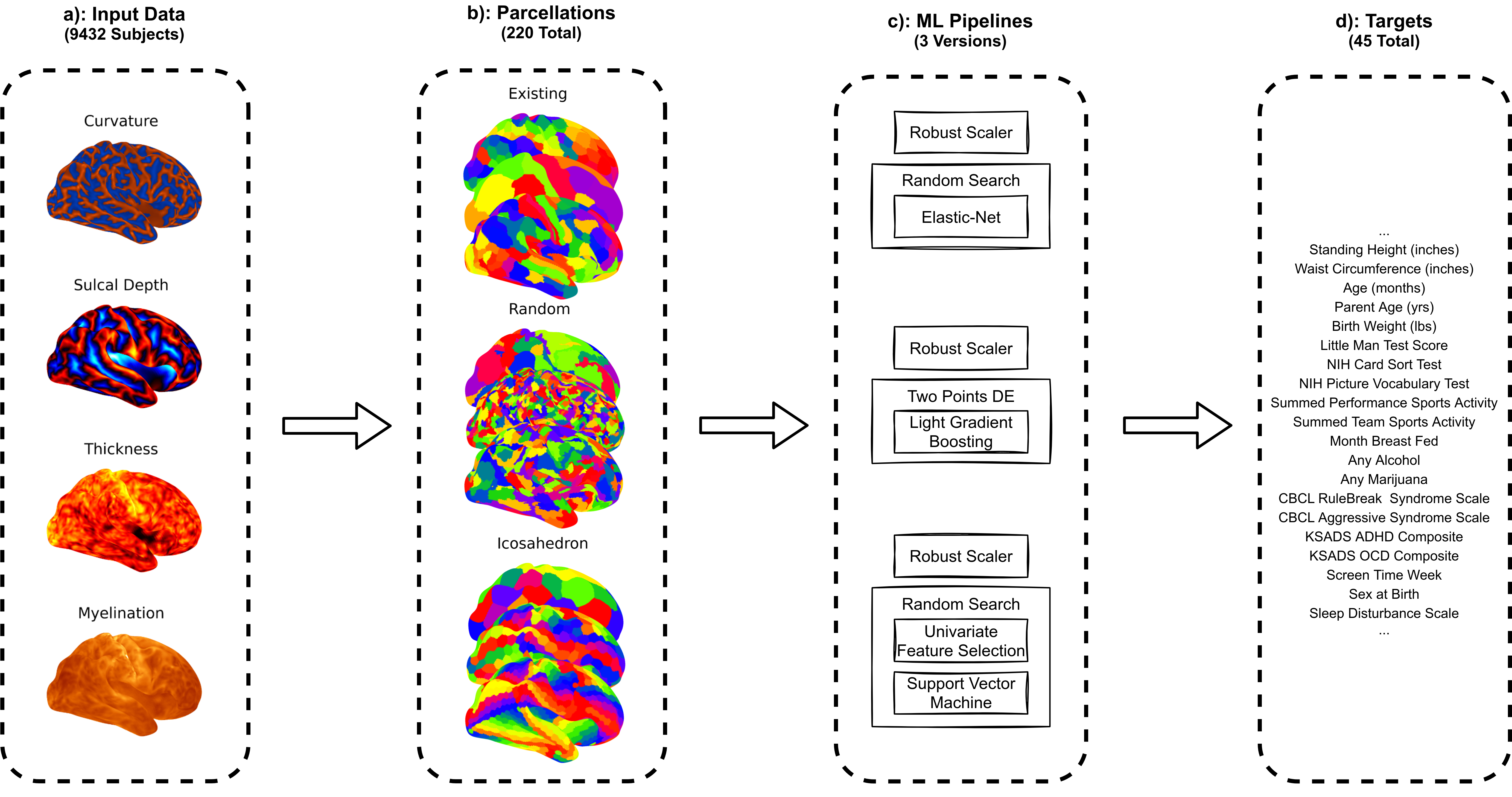
The base experiment conducted within this project was a systematic test of performance of different pre-defined parcellations. The structure of the evaluation is shown below:
-
a). This study uses baseline data from the ABCD Study NDA Collection 3165 Release. Specifically, we concatenate 9,432 participant’s structural MRI measures to use as input features for ML (See Input Data for more information).
-
b). We test a mix of mostly random and existing parcellations (See Parcellations).
-
c). Three different ML pipelines are used, each based on a different popular base estimator (See ML Pipelines).
-
d). In total we employ 45 different phenotypic target variables (See Target Variables).
We evaluated each combination of target variable, parcellation and ML pipeline with five-fold cross validation using the full set of available participants. The CV fold structure was kept constant and therefore directly comparable across all combinations of ML pipeline, target variable and parcellation. This evaluation procedure was used to generate different metrics of performance, R2 for regression predictors and area under the receiver operator characteristic curve (ROC AUC) for binary predictors, for each of the combinations. Performance metrics were then converted in the results into a measure of Mean Rank.
Base Experiment Results
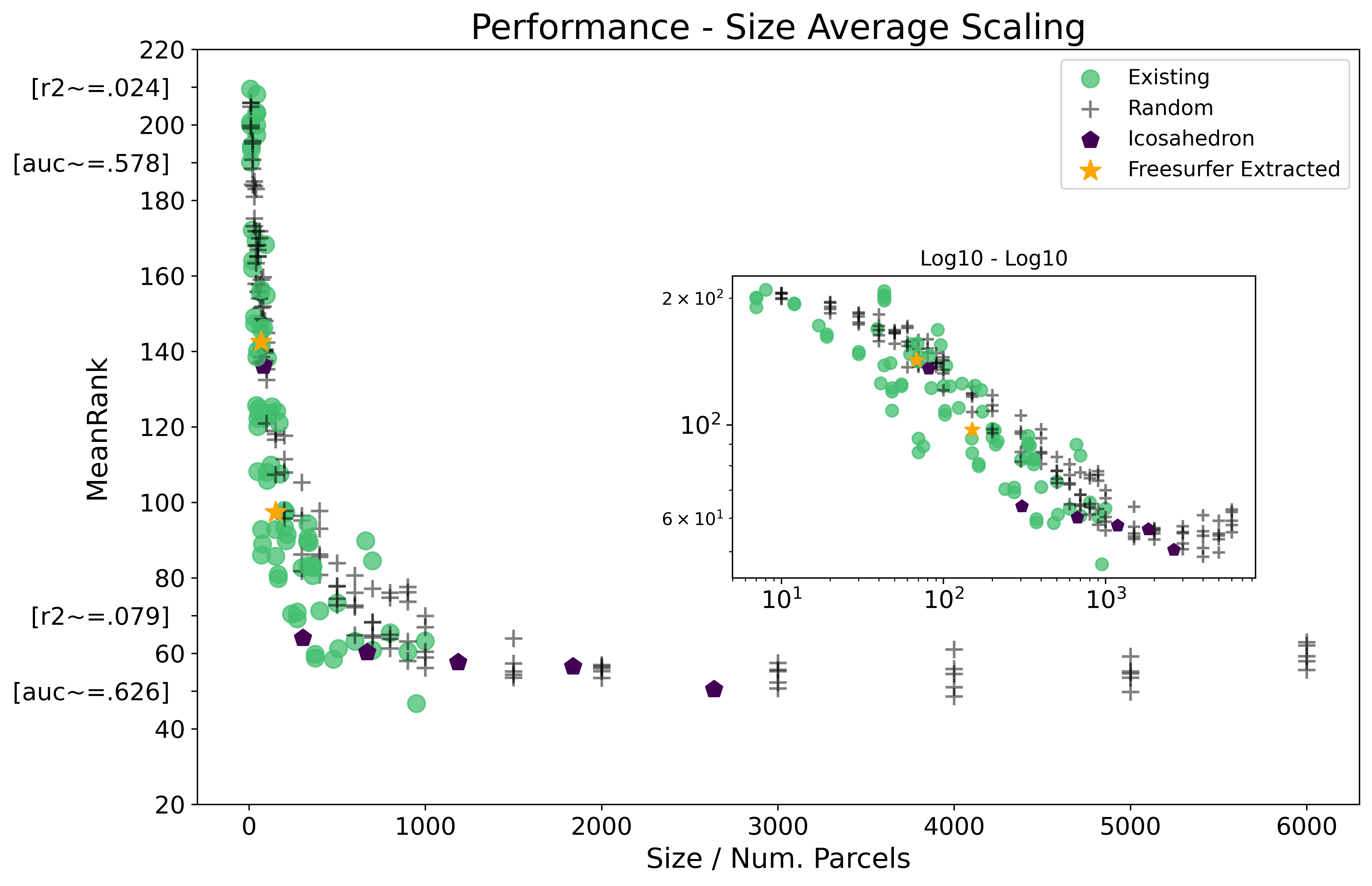
The below figure plots performance, as measured by mean relative ranking between all 220 parcellations, against the number of parcels / size in each parcellation. Results are further colored by type of parcellation and a log10-log10 inset of the same plot is provided. It may be useful to also review the Intro to Results page first, which provides a gradual introduction to the format the results are plotted with below.
Click the figure above to open an interactive version of the plot
-
There is a relationship between performance, as measured by mean relative ranking, and parcellation size. Up to ~Size 4000 a power law relationship appears to hold, with scaling exponent .-2753. See results table.
-
Existing parcellations outperformed randomly generated parcellations when controlling for the influence of size, however existing parcellations tended to have fewer parcels than our results suggest to be most predictive (See Results by Parcellation Type).
-
The general pattern was stable across ML Pipelines, but when compared inter-pipeline, the SVM based pipeline was most competitive. See Results by Pipeline.
-
How stable are these results across different target variables? See Results by Target.
Multiple Parcellation Strategies
As an additional set of analyses we sought to characterize the potential gains in performance from employing strategies that can make use of information from multiple parcellations in order to inform predictions. These extensions to the base analysis can be broken up into three different types: choice of parcellation as a nested hyper-parameter - (“Grid”), ensembling over multiple parcellations using voting - (“Voted”), and ensembling using stacking - (“Stacked”). See Multiple Parcellations Setup for more detailed information on how this experiment was structured.
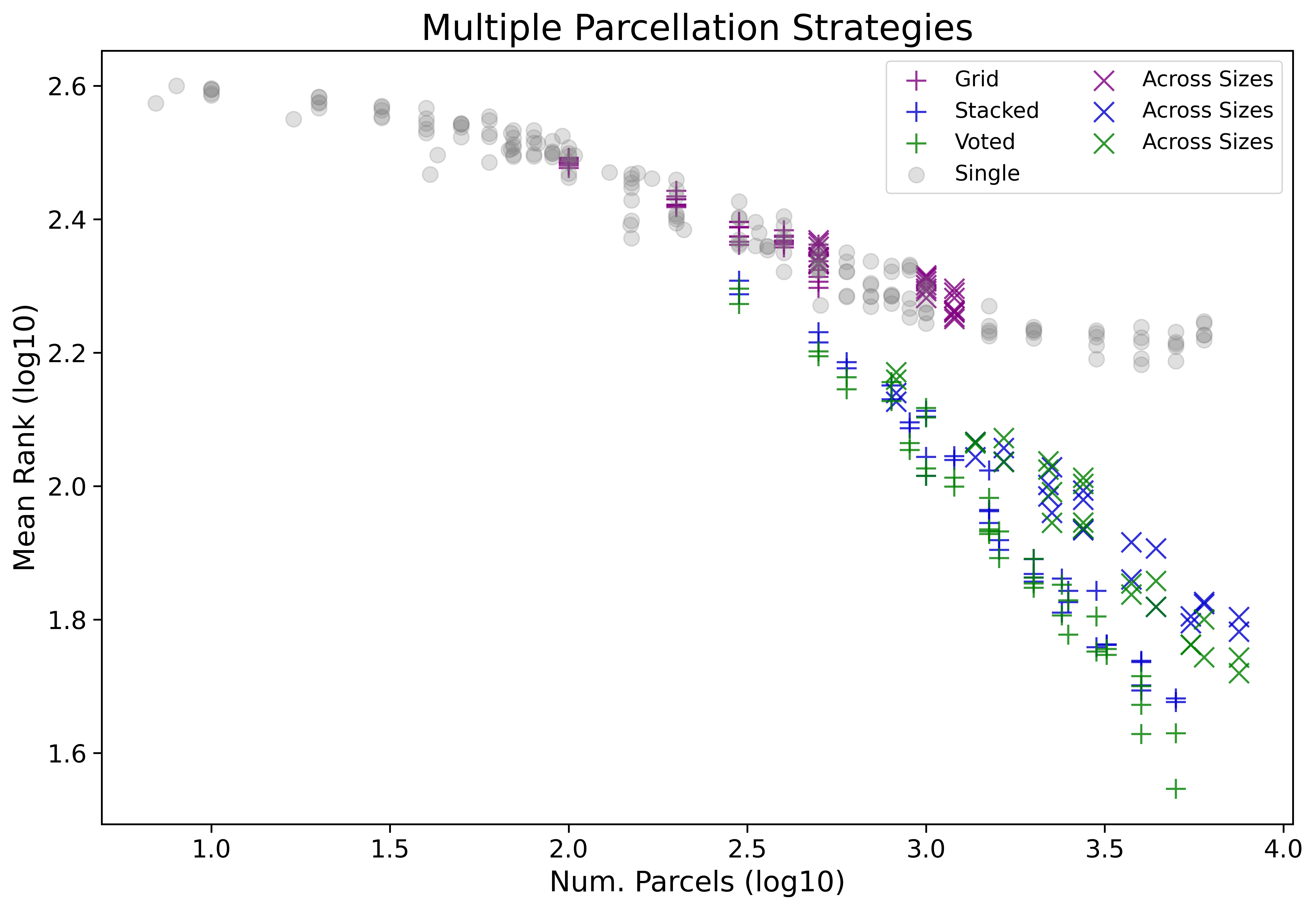
The figure below compares the prior single parcellation only results to the introduced multiple parcellation strategies. The plotted mean ranks are therefore computed now between 412 (220 single parcellation and 192 multiple parcellation based) configurations. The results are further broken down by if the pool of parcellations was sourced from fixed sizes or across multiple sizes (See Multiple Parcellations Evaluation).
 Click the figure above to open an interactive version of the plot
Click the figure above to open an interactive version of the plot
-
Ensemble methods across multiple parcellations outperform single parcellation based methods (See Single vs. Ensembled Parcellations).
-
We did not observe a large difference between ensemble strategies Voted and Stacked - unless we breakdown results by binary vs. regression!
-
Sourcing random parcellation for ensembling from fixed sizes outperforms sourcing parcellations from a range of sizes (See Fixed vs. Across Sizes).
-
The SVM based ensembles were once again better than the others, with the exception now of the special ‘All’ ensemble. See Ensemble Results by Pipeline.
-
Searching over multiple parcellations as hyper-parameter was not a successful strategy. See Grid vs. Random.
-
All options considered, what are the best strategies across different sizes?
Discussion Points
-
There is certainly a relationship between parcellation scale and performance, but what factors influence this relationship?
-
Why do we see a performance boost from increasing parcellation resolution?
-
This project makes a lot of different comparisons, with this in mind we provide some practical recommendations for researchers.
-
Notably, performance may not in practice be the only metric of interest, instead there are a number of Performance Trade-Offs to consider, e.g., runtime and downstream interpretation complexity.
-
This project required an incredible amount of computations, but we also made a great deal of effort to optimize performance wherever possible.
-
There are a number of possible extensions / future work.
Conclusion
In testing a variety of parcellation schemes and ML modeling approaches, we have identified an apparent power law scaling of increasing predictive performance by increasing parcellation resolution. The details of this relationship were found to vary according to type of parcellation as well as ML pipeline employed, though the general pattern proved stable. The large sample size, range of predictive targets, and collection of existing and random parcellations tested all serve to lend confidence to the observed results. Researchers selecting a parcellation for predictive modelling may wish to consider this size-performance trade-off in addition to other factors such as interpretability and computational resources. We also highlighted important factors that improved performance above and beyond the size-scaling, for example, finding existing parcellations performed better than randomly generated parcellations. Further, we demonstrated the benefit of ensembling over multiple parcellations, which yielded a performance boost relative to results from single parcellations.
Authors
Sage Hahn, Max M. Owens, DeKang Yuan, Anthony C Juliano, Alexandra Potter, Hugh Garavan, Nicholas Allgaier
Departments of Complex Systems and Psychiatry, University of Vermont, Burlington, VT 05401
Acknowledgments
-
Sage Hahn, Max M. Owens, DeKang Yuan and Anthony C Juliano were supported by NIDA grant T32DA043593
-
Data used in the preparation of this article were obtained from the ABCD Study held in the NDA. This is a multisite, longitudinal study designed to recruit more than 10,000 children ages 9–10 years old and follow them over 10 years into early adulthood. The ABCD study is supported by the National Institutes of Health and additional federal partners under award numbers U01DA041048, U01DA050989, U01DA051016, U01DA041022, U01DA051018, U01DA051037, U01DA050987, U01DA041174, U01DA041106, U01DA041117, U01DA041028, U01DA041134, U01DA050988, U01DA051039, U01DA041156, U01DA041025, U01DA041120, U01DA051038, U01DA041148, U01DA041093, U01DA041089, U24DA041123 and U24DA041147. A full list of supporters is available at https://abcdstudy.org/federal-partners.html. A listing of participating sites and a complete listing of the study investigators can be found at https://abcdstudy.org/consortium_members/.
-
Computations were performed on the Vermont Advanced Computing Core supported, in part, by NSF award number OAC-1827314.
-
We would also like to thank the other members of the Hugh Garavan lab for their support throughout this project.



Site Map
This project website is surprisingly expansive when considering nested hyper-links. Listed below are links in alphabetical order to all main site pages (many of which include multiple sub-sections):
- All Results by Target Table Results
- All Raw Results by Pipeline
- All Targets
- Base Results Size Differences
- Base Results
- Base Results Extra
- Effects of Feature Selection
- Ensemble Comparison
- Ensemble Inter-Pipe Table Results
- Ensemble Intra-Pipe Table Results
- Ensemble Raw Intra-Pipeline Comparison
- Ensemble Results by Pipeline
- Estimate Powerlaw
- Evaluation Structure
- Full Table Results
- Future Work
- Grid vs. Random
- Input Data
- Inter-Pipeline Table Results
- Interactive Figure 1
- Interactive Figure 1 R2
- Interactive Figure 1 ROC AUC
- Interactive Figure 2
- Interactive Figure 2 Base
- Interactive Figure 2 R2
- Interactive Figure 2 Base R2
- Interactive Figure 2 ROC AUC
- Interactive Figure 2 Base ROC AUC
- Interactive Figure 3
- Interactive Figure 3 R2
- Interactive Figure 3 ROC AUC
- Interactive Figure 4
- Interactive Figure 4 R2
- Interactive Figure 4 ROC AUC
- Interactive Figure 5
- Interactive Figure 5 Base
- Interactive Figure 5 R2
- Interactive Figure 5 Base R2
- Interactive Figure 5 ROC AUC
- Interactive Figure 5 Base ROC AUC
- Interactive Figure 6
- Interactive Figure 6 Base
- Interactive Figure 6 R2
- Interactive Figure 6 Base R2
- Interactive Figure 6 ROC AUC
- Interactive Figure 6 Base ROC AUC
- Interactive Figure 7
- Interactive Figure 7 R2
- Interactive Figure 7 ROC AUC
- Intra-Pipeline Table Results
- Intro to Results
- ML Pipelines
- ML for Neuroimaging
- Multiple Parcellation Strategies Setup
- Outliers
- Parcellations Viz
- Parcellations
- Performance Optimizations
- Performance Scaling
- Raw Intra-Pipeline Comparison
- Recommendations
- Resampling Parcellations
- Results by Clusters of Targets
- Results by Pipeline
- Results by Pipeline Median
- Results by Target Table Results
- Results by Target
- Scaling Exponent
- Scaling Issues
- Single vs. Ensembled Parcellations
- Special Ensembles
- Target Variables
- Trade-Offs
- Website Info
- Whats best?
- Why a performance boost?
- Why ensemble boost?